Here, we’ll go over some examples of using ITT. First we need to load the library before getting in to some sample use cases.
ITT With 5 bootstrap samples
options <- SEQopts(# tells SEQuential to create Kaplan-Meier curves
km.curves = TRUE,
# tells SEQuential to bootstrap
bootstrap = TRUE,
# tells SEQuential to run bootstraps 5 times
bootstrap.nboot = 5)
# use example data
data <- SEQdata
model <- SEQuential(data, id.col = "ID",
time.col = "time",
eligible.col = "eligible",
treatment.col = "tx_init",
outcome.col = "outcome",
time_varying.cols = c("N", "L", "P"),
fixed.cols = "sex",
method = "ITT",
options = options)
#> Non-required columns provided, pruning for efficiency
#> Pruned
#> Expanding Data...
#> Expansion Successful
#> Moving forward with ITT analysis
#> Bootstrapping with 80 % of data 5 times
#> ITT model created successfully
#> Creating Survival curves
#> Completed
km_curve(model, plot.type = "risk") # retrieve risk plot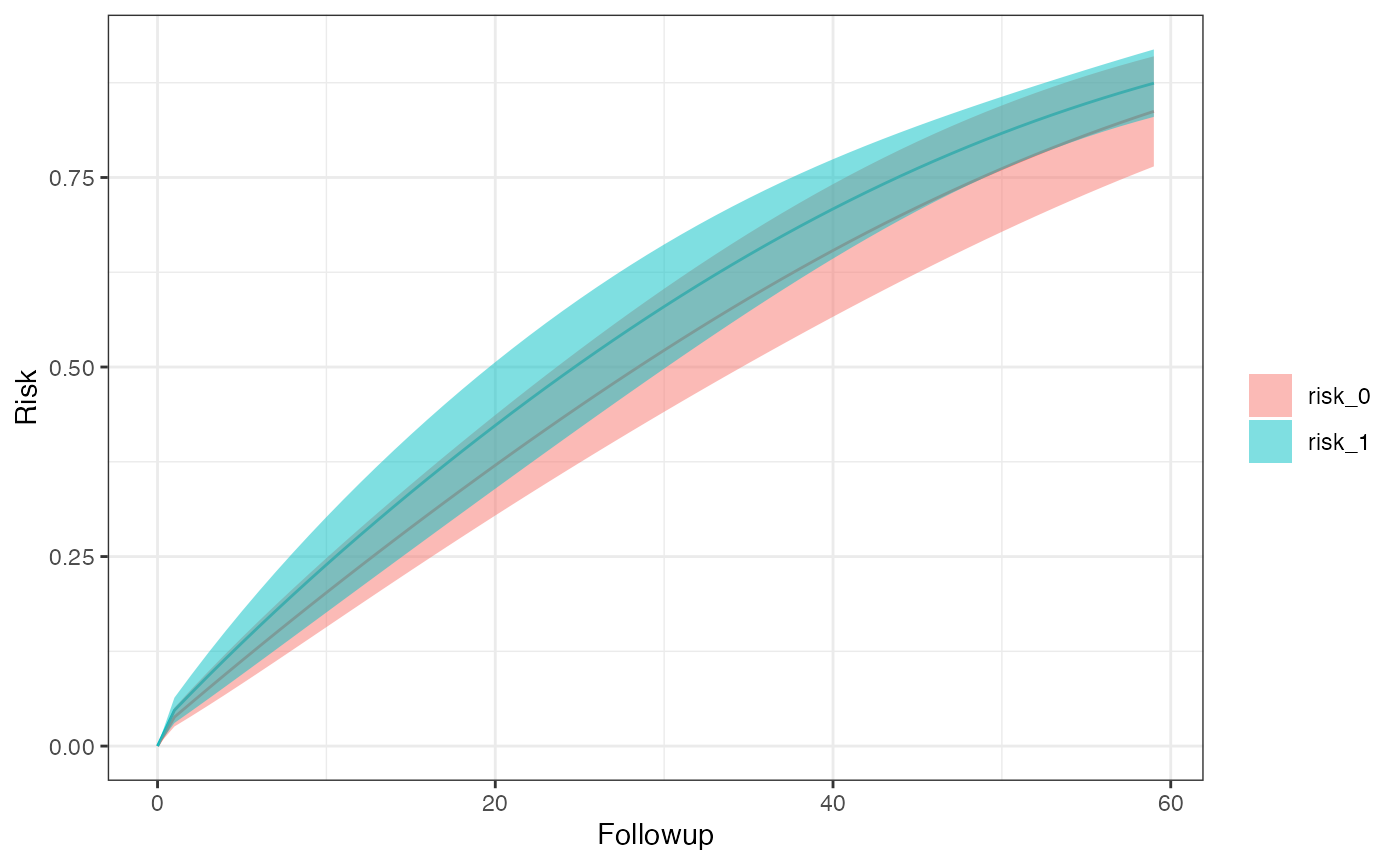
risk_data(model)
#> Method A Risk 95% LCI 95% UCI SE
#> <char> <char> <num> <num> <num> <num>
#> 1: ITT 0 0.8372582 0.7646336 0.9098829 0.03705407
#> 2: ITT 1 0.8744359 0.8299662 0.9189056 0.02268902
risk_comparison(model)
#> A_x A_y Risk Ratio RR 95% LCI RR 95% UCI Risk Differerence RD 95% LCI
#> <fctr> <fctr> <num> <num> <num> <num> <num>
#> 1: risk_0 risk_1 1.0444041 0.9685600 1.126187 0.03717768 -0.02814011
#> 2: risk_1 risk_0 0.9574838 0.8879519 1.032461 -0.03717768 -0.10249547
#> RD 95% UCI
#> <num>
#> 1: 0.10249547
#> 2: 0.02814011ITT with 5 bootstrap samples and losses-to-followup
options <- SEQopts(km.curves = TRUE,
bootstrap = TRUE,
bootstrap.nboot = 5,
# tells SEQuential to expect LTFU as the censoring column
cense = "LTFU",
# tells SEQuential to treat this column as the
# censoring eligibility column
cense.eligible = "eligible_cense")
# use example data for LTFU
data <- SEQdata.LTFU
model <- SEQuential(data, id.col = "ID",
time.col = "time",
eligible.col = "eligible",
treatment.col = "tx_init",
outcome.col = "outcome",
time_varying.cols = c("N", "L", "P"),
fixed.cols = "sex",
method = "ITT",
options = options)
#> Non-required columns provided, pruning for efficiency
#> Pruned
#> Expanding Data...
#> Expansion Successful
#> Moving forward with ITT analysis
#> Bootstrapping with 80 % of data 5 times
#> ITT model created successfully
#> Creating Survival curves
#> Completed
km_curve(model, plot.type = "risk")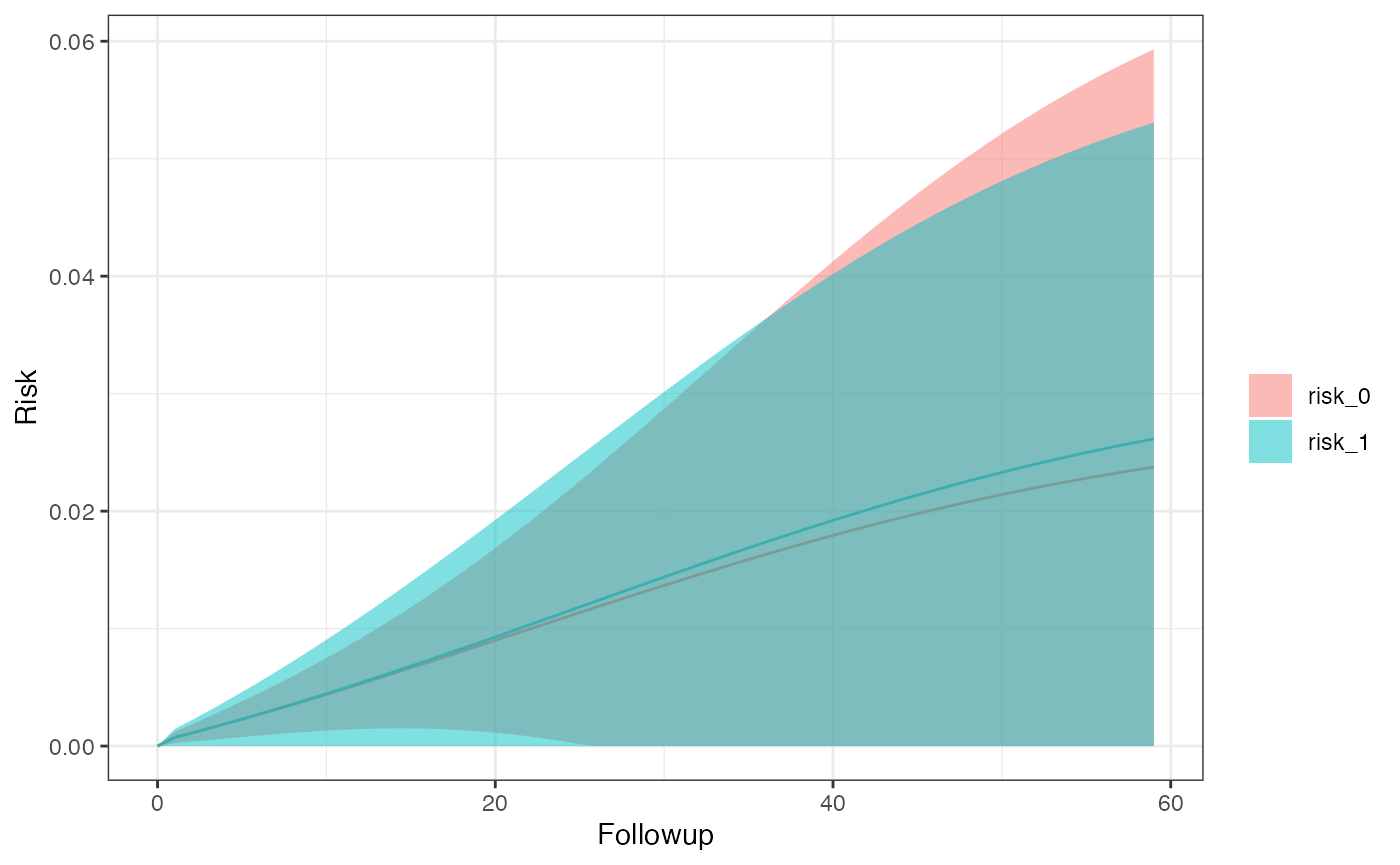
risk_data(model)
#> Method A Risk 95% LCI 95% UCI SE
#> <char> <char> <num> <num> <num> <num>
#> 1: ITT 0 0.02374360 0 0.05930116 0.01814195
#> 2: ITT 1 0.02614576 0 0.05307095 0.01373759
risk_comparison(model)
#> A_x A_y Risk Ratio RR 95% LCI RR 95% UCI Risk Differerence
#> <fctr> <fctr> <num> <num> <num> <num>
#> 1: risk_0 risk_1 1.1011710 0.5047356 2.402402 0.002402164
#> 2: risk_1 risk_0 0.9081242 0.4162501 1.981235 -0.002402164
#> RD 95% LCI RD 95% UCI
#> <num> <num>
#> 1: -0.009778817 0.014583145
#> 2: -0.014583145 0.009778817ITT with 5 bootstrap samples and competing events
options <- SEQopts(km.curves = TRUE,
bootstrap = TRUE,
bootstrap.nboot = 5,
# Using LTFU as our competing event
compevent = "LTFU")
data <- SEQdata.LTFU
model <- SEQuential(data, id.col = "ID",
time.col = "time",
eligible.col = "eligible",
treatment.col = "tx_init",
outcome.col = "outcome",
time_varying.cols = c("N", "L", "P"),
fixed.cols = "sex",
method = "ITT",
options = options)
#> Non-required columns provided, pruning for efficiency
#> Pruned
#> Expanding Data...
#> Expansion Successful
#> Moving forward with ITT analysis
#> Bootstrapping with 80 % of data 5 times
#> ITT model created successfully
#> Creating Survival curves
#> Completed
km_curve(model, plot.type = "risk")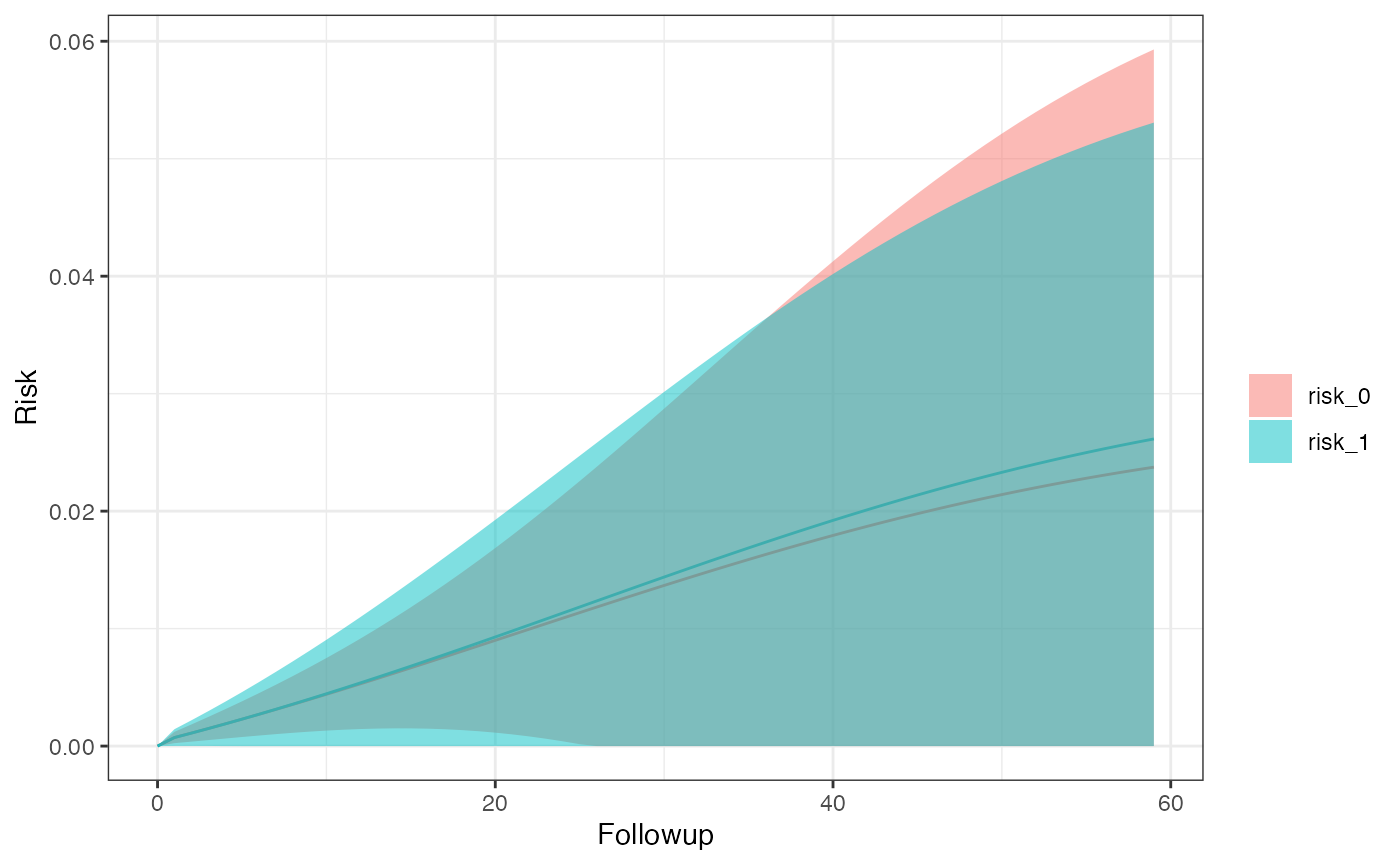
risk_data(model)
#> Method A Risk 95% LCI 95% UCI SE
#> <char> <char> <num> <num> <num> <num>
#> 1: ITT 0 0.02185652 0 0.05286838 0.01582267
#> 2: ITT 1 0.02381601 0 0.04835280 0.01251900
risk_comparison(model)
#> A_x A_y Risk Ratio RR 95% LCI RR 95% UCI Risk Differerence
#> <fctr> <fctr> <num> <num> <num> <num>
#> 1: inc_0 inc_1 1.0896524 0.5100805 2.327755 0.001959489
#> 2: inc_1 inc_0 0.9177239 0.4295985 1.960475 -0.001959489
#> RD 95% LCI RD 95% UCI
#> <num> <num>
#> 1: -0.007920832 0.011839810
#> 2: -0.011839810 0.007920832ITT hazard ratio with 5 bootstrap samples and competing events
options <- SEQopts(# km.curves must be set to FALSE to turn on hazard
# ratio creation
km.curves = FALSE,
# set hazard to TRUE for hazard ratio creation
hazard = TRUE,
bootstrap = TRUE,
bootstrap.nboot = 5,
compevent = "LTFU")
data <- SEQdata.LTFU
model <- SEQuential(data, id.col = "ID",
time.col = "time",
eligible.col = "eligible",
treatment.col = "tx_init",
outcome.col = "outcome",
time_varying.cols = c("N", "L", "P"),
fixed.cols = "sex",
method = "ITT",
options = options)
#> Non-required columns provided, pruning for efficiency
#> Pruned
#> Expanding Data...
#> Expansion Successful
#> Moving forward with ITT analysis
#> Bootstrapping with 80 % of data 5 times
#> Completed
# retrieve hazard ratios
hazard_ratio(model)
#> Hazard ratio LCI UCI
#> 1.0033697 0.5993046 1.6798651ITT with 5 bootstrap samples and competing events in subgroups defined by sex
options <- SEQopts(km.curves = TRUE,
bootstrap = TRUE,
bootstrap.nboot = 5,
compevent = "LTFU",
# define the subgroup
subgroup = "sex")
data <- SEQdata.LTFU
model <- SEQuential(data, id.col = "ID",
time.col = "time",
eligible.col = "eligible",
treatment.col = "tx_init",
outcome.col = "outcome",
time_varying.cols = c("N", "L", "P"),
fixed.cols = "sex",
method = "ITT",
options = options)
#> Non-required columns provided, pruning for efficiency
#> Pruned
#> Expanding Data...
#> Expansion Successful
#> Moving forward with ITT analysis
#> Bootstrapping with 80 % of data 5 times
#> ITT model created successfully
#> Creating Survival Curves for sex_0
#> Creating Survival Curves for sex_1
#> Completed
km_curve(model, plot.type = "risk")
#> $sex_0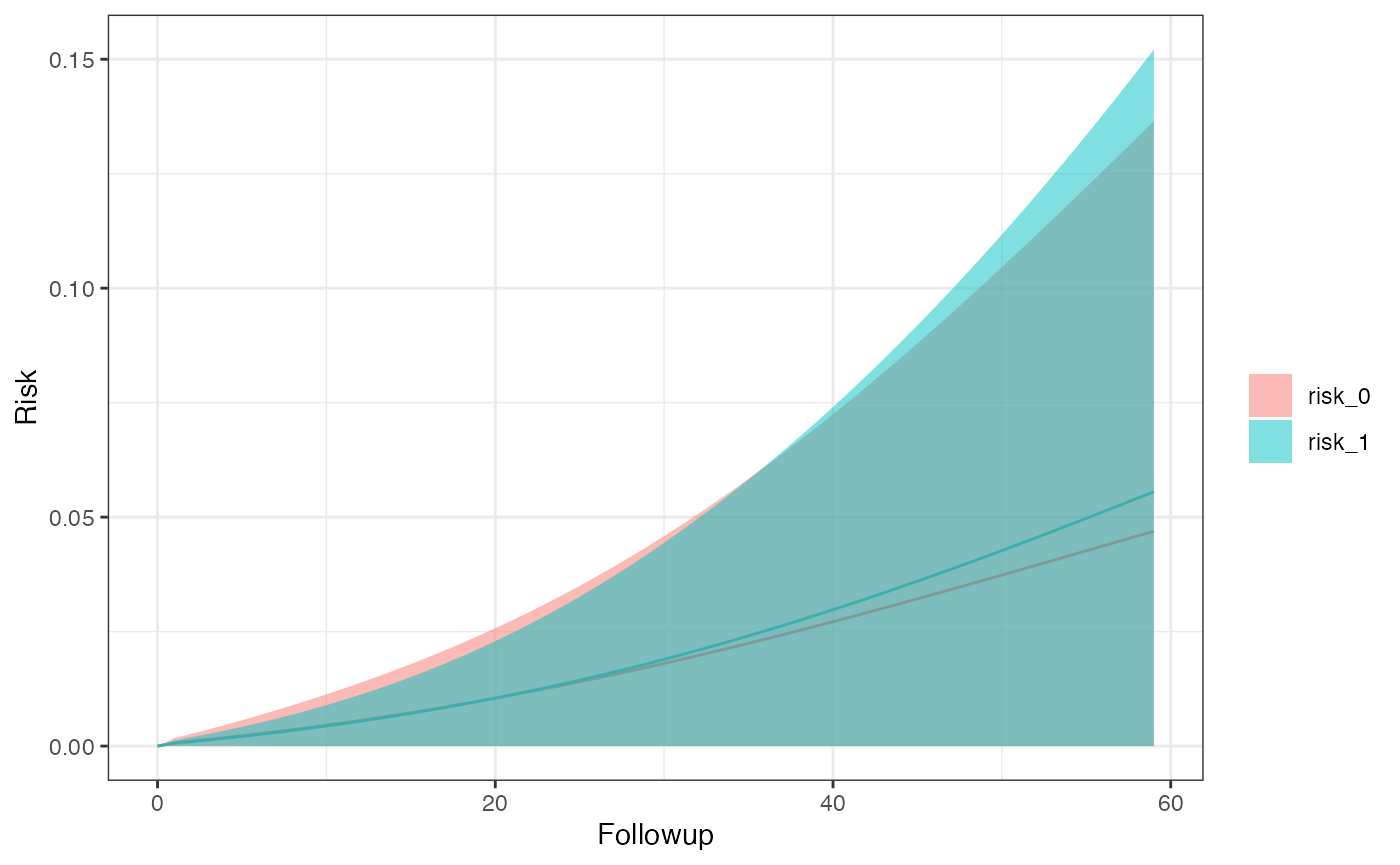
#>
#> $sex_1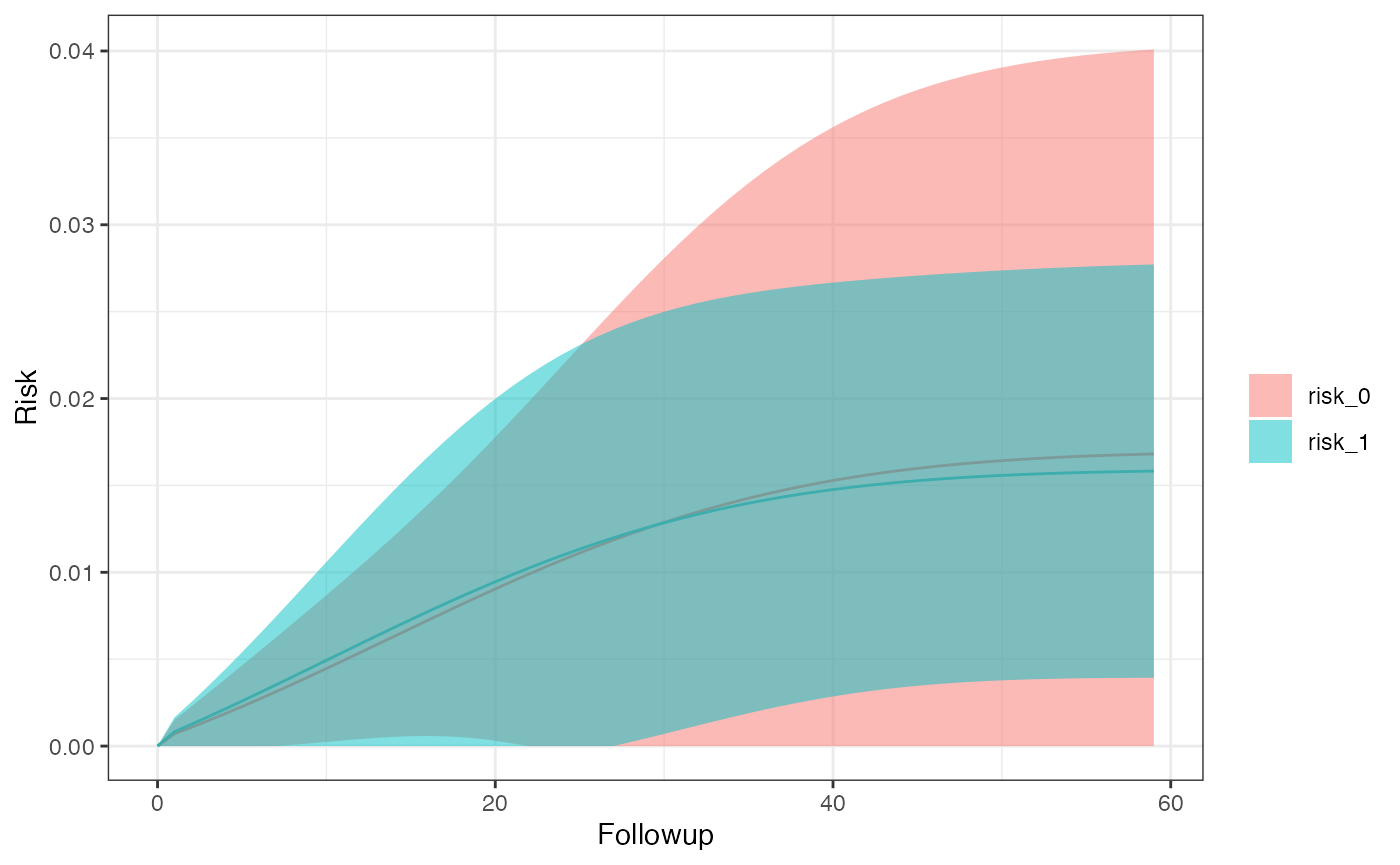
risk_data(model)
#> $sex_0
#> Method A Risk 95% LCI 95% UCI SE
#> <char> <char> <num> <num> <num> <num>
#> 1: ITT 0 0.04213833 0 0.1185452 0.03898383
#> 2: ITT 1 0.04911213 0 0.1322803 0.04243352
#>
#> $sex_1
#> Method A Risk 95% LCI 95% UCI SE
#> <char> <char> <num> <num> <num> <num>
#> 1: ITT 0 0.01577026 0.000000000 0.03706622 0.010865485
#> 2: ITT 1 0.01484521 0.003496358 0.02619406 0.005790336
risk_comparison(model)
#> $sex_0
#> A_x A_y Risk Ratio RR 95% LCI RR 95% UCI Risk Differerence
#> <fctr> <fctr> <num> <num> <num> <num>
#> 1: inc_0 inc_1 1.1654977 1.876650e-08 72383489 0.006973797
#> 2: inc_1 inc_0 0.8580026 1.381531e-08 53286436 -0.006973797
#> RD 95% LCI RD 95% UCI
#> <num> <num>
#> 1: -0.02096462 0.03491222
#> 2: -0.03491222 0.02096462
#>
#> $sex_1
#> A_x A_y Risk Ratio RR 95% LCI RR 95% UCI Risk Differerence RD 95% LCI
#> <fctr> <fctr> <num> <num> <num> <num> <num>
#> 1: inc_0 inc_1 0.9413422 0.4013911 2.207635 -0.0009250492 -0.01668972
#> 2: inc_1 inc_0 1.0623130 0.4529734 2.491336 0.0009250492 -0.01483962
#> RD 95% UCI
#> <num>
#> 1: 0.01483962
#> 2: 0.01668972