Here, we’ll go over some examples of using dose-response. First we need to load the library before getting in to some sample use cases.
Currently, dose-response analysis through SEQuential only supports binary treatment values. Therefore; running multinomial models will lead to errors.
Dose-response With 5 bootstrap samples
options <- SEQopts(# tells SEQuential to create Kaplan-Meier curves
km.curves = TRUE,
# tells SEQuential to bootstrap
bootstrap = TRUE,
# tells SEQuential to run bootstraps 5 times
bootstrap.nboot = 5)
# use example data
data <- SEQdata
model <- SEQuential(data, id.col = "ID",
time.col = "time",
eligible.col = "eligible",
treatment.col = "tx_init",
outcome.col = "outcome",
time_varying.cols = c("N", "L", "P"),
fixed.cols = "sex",
method = "dose-response",
options = options)
#> Non-required columns provided, pruning for efficiency
#> Pruned
#> Expanding Data...
#> Expansion Successful
#> Moving forward with dose-response analysis
#> Bootstrapping with 80 % of data 5 times
#> dose-response model created successfully
#> Creating Survival curves
#> Completed
km_curve(model, plot.type = "risk") # retrieve risk plot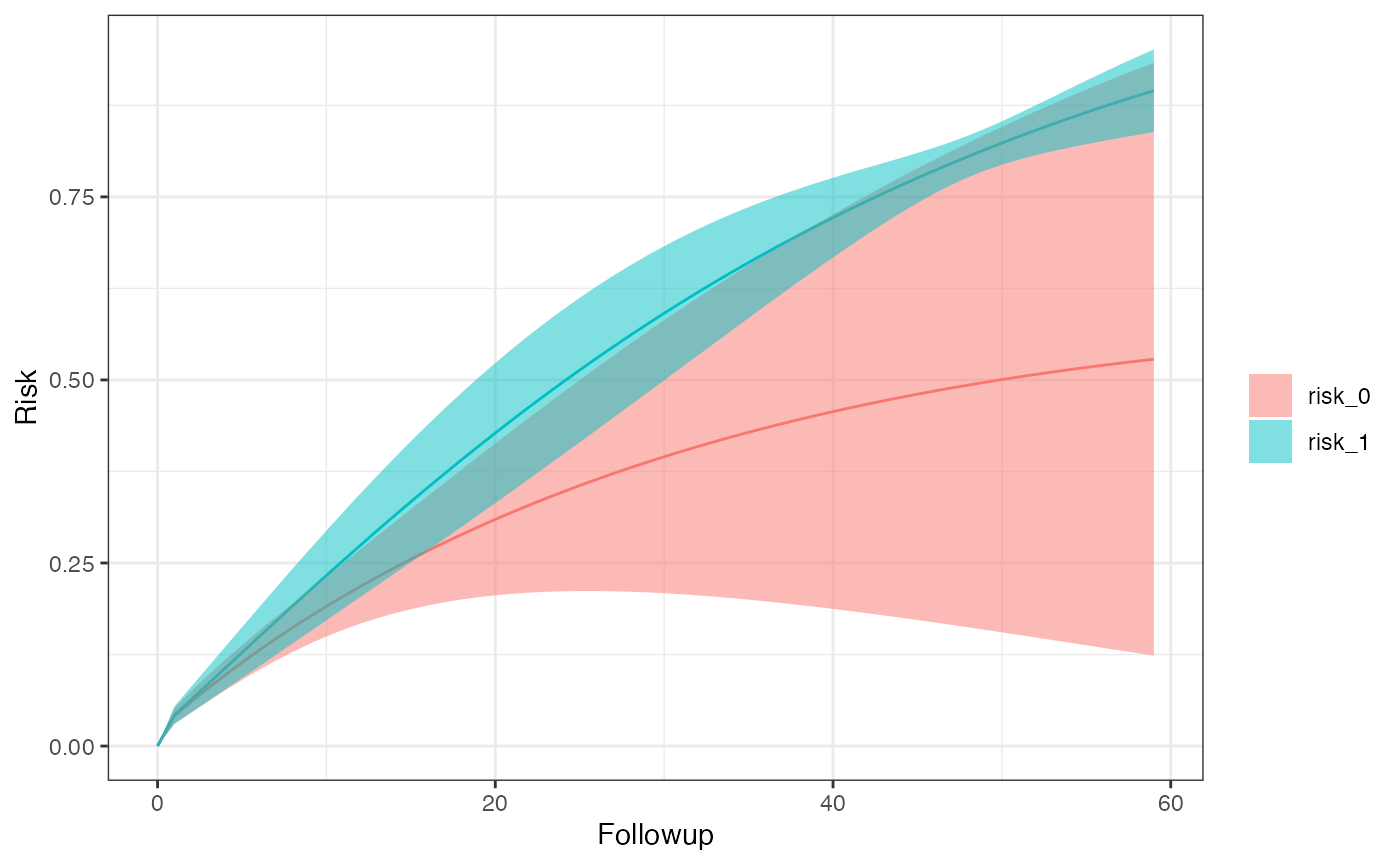
risk_data(model)
#> Method A Risk 95% LCI 95% UCI SE
#> <char> <char> <num> <num> <num> <num>
#> 1: dose-response 0 0.5282782 0.1234894 0.9330671 0.20652872
#> 2: dose-response 1 0.8949096 0.8385037 0.9513155 0.02877903
risk_comparison(model)
#> A_x A_y Risk Ratio RR 95% LCI RR 95% UCI Risk Differerence RD 95% LCI
#> <fctr> <fctr> <num> <num> <num> <num> <num>
#> 1: risk_0 risk_1 1.6940120 0.7764767 3.695766 0.3666314 -0.07309331
#> 2: risk_1 risk_0 0.5903146 0.2705799 1.287869 -0.3666314 -0.80635612
#> RD 95% UCI
#> <num>
#> 1: 0.80635612
#> 2: 0.07309331Dose-response with 5 bootstrap samples and losses-to-followup
options <- SEQopts(km.curves = TRUE,
bootstrap = TRUE,
bootstrap.nboot = 5,
# tells SEQuential to expect LTFU as the censoring column
cense = "LTFU",
# tells SEQuential to treat this column as the
# censoring eligibility column
cense.eligible = "eligible_cense")
# use example data for LTFU
data <- SEQdata.LTFU
model <- SEQuential(data, id.col = "ID",
time.col = "time",
eligible.col = "eligible",
treatment.col = "tx_init",
outcome.col = "outcome",
time_varying.cols = c("N", "L", "P"),
fixed.cols = "sex",
method = "dose-response",
options = options)
#> Non-required columns provided, pruning for efficiency
#> Pruned
#> Expanding Data...
#> Expansion Successful
#> Moving forward with dose-response analysis
#> Bootstrapping with 80 % of data 5 times
#> dose-response model created successfully
#> Creating Survival curves
#> Completed
km_curve(model, plot.type = "risk")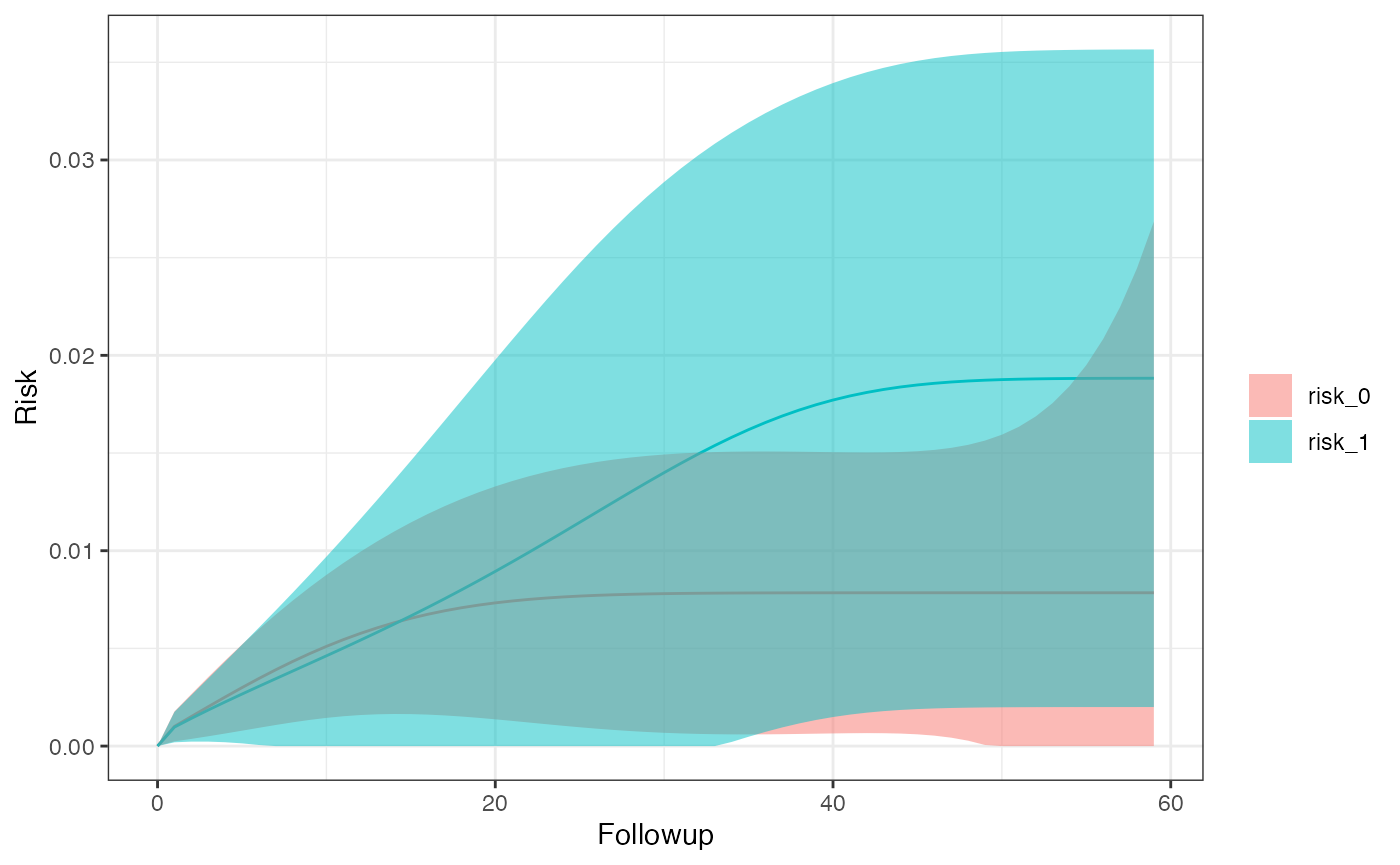
risk_data(model)
#> Method A Risk 95% LCI 95% UCI SE
#> <char> <char> <num> <num> <num> <num>
#> 1: dose-response 0 0.007847443 0.000000000 0.02684988 0.009695299
#> 2: dose-response 1 0.018827788 0.001997953 0.03565762 0.008586808
risk_comparison(model)
#> A_x A_y Risk Ratio RR 95% LCI RR 95% UCI Risk Differerence RD 95% LCI
#> <fctr> <fctr> <num> <num> <num> <num> <num>
#> 1: risk_0 risk_1 2.3992259 0.44856850 12.832566 0.01098034 -0.01349983
#> 2: risk_1 risk_0 0.4168011 0.07792674 2.229314 -0.01098034 -0.03546052
#> RD 95% UCI
#> <num>
#> 1: 0.03546052
#> 2: 0.01349983Dose-response with 5 bootstrap samples and competing events
options <- SEQopts(km.curves = TRUE,
bootstrap = TRUE,
bootstrap.nboot = 5,
# Using LTFU as our competing event
compevent = "LTFU")
data <- SEQdata.LTFU
model <- SEQuential(data, id.col = "ID",
time.col = "time",
eligible.col = "eligible",
treatment.col = "tx_init",
outcome.col = "outcome",
time_varying.cols = c("N", "L", "P"),
fixed.cols = "sex",
method = "dose-response",
options = options)
#> Non-required columns provided, pruning for efficiency
#> Pruned
#> Expanding Data...
#> Expansion Successful
#> Moving forward with dose-response analysis
#> Bootstrapping with 80 % of data 5 times
#> dose-response model created successfully
#> Creating Survival curves
#> Completed
km_curve(model, plot.type = "risk")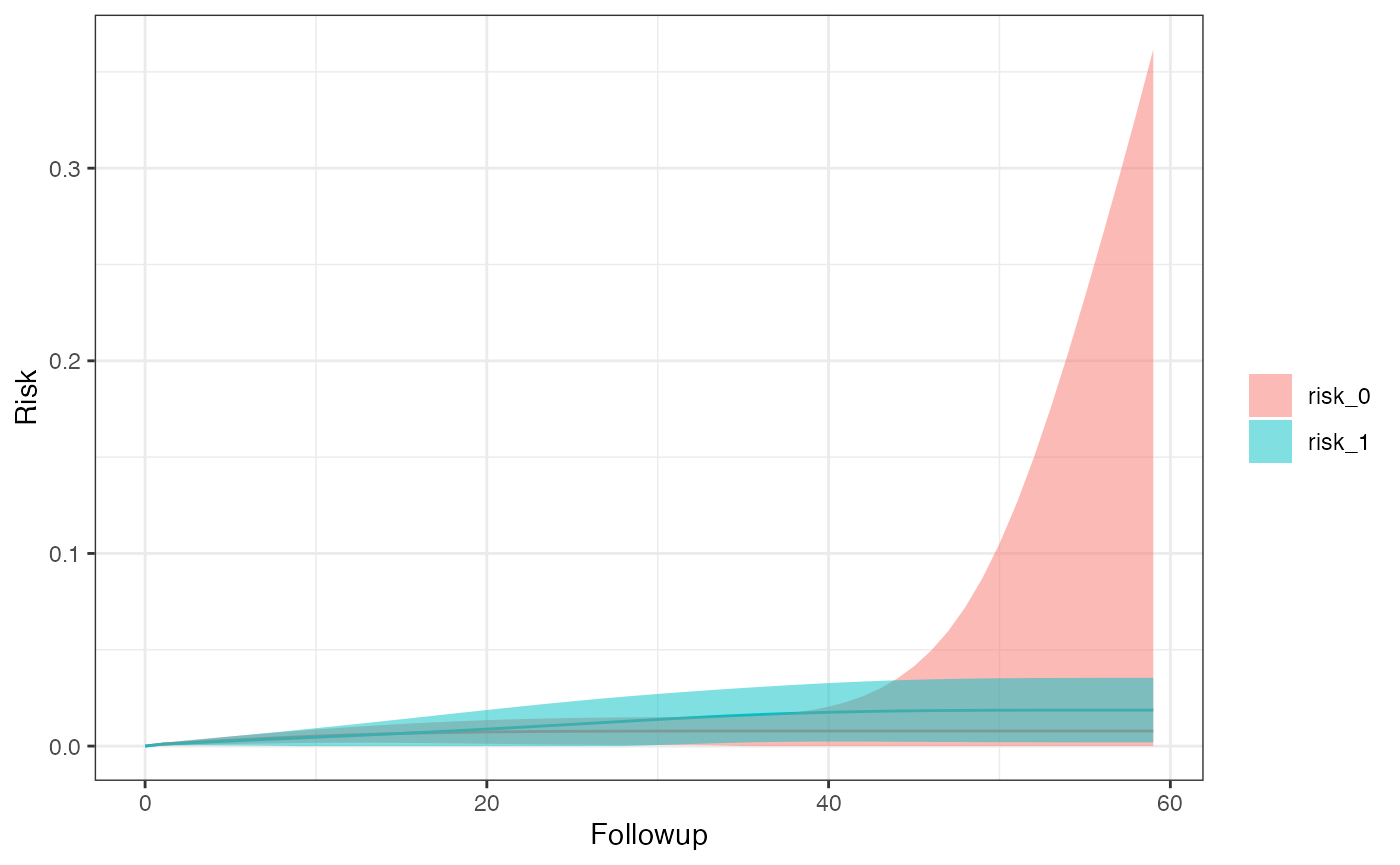
risk_data(model)
#> Method A Risk 95% LCI 95% UCI SE
#> <char> <char> <num> <num> <num> <num>
#> 1: dose-response 0 0.007586789 0 0.25972299 0.12864328
#> 2: dose-response 1 0.003641046 0 0.02703035 0.01193354
risk_comparison(model)
#> A_x A_y Risk Ratio RR 95% LCI RR 95% UCI Risk Differerence RD 95% LCI
#> <fctr> <fctr> <num> <num> <num> <num> <num>
#> 1: inc_0 inc_1 0.4799192 0.02752717 8.367096 -0.003945743 -0.2462717
#> 2: inc_1 inc_0 2.0836841 0.11951578 36.327751 0.003945743 -0.2383802
#> RD 95% UCI
#> <num>
#> 1: 0.2383802
#> 2: 0.2462717Dose-response hazard ratio with 5 bootstrap samples and competing events
options <- SEQopts(# km.curves must be set to FALSE to turn on hazard
# ratio creation
km.curves = FALSE,
# set hazard to TRUE for hazard ratio creation
hazard = TRUE,
bootstrap = TRUE,
bootstrap.nboot = 5,
compevent = "LTFU")
data <- SEQdata.LTFU
model <- SEQuential(data, id.col = "ID",
time.col = "time",
eligible.col = "eligible",
treatment.col = "tx_init",
outcome.col = "outcome",
time_varying.cols = c("N", "L", "P"),
fixed.cols = "sex",
method = "dose-response",
options = options)
#> Non-required columns provided, pruning for efficiency
#> Pruned
#> Expanding Data...
#> Expansion Successful
#> Moving forward with dose-response analysis
#> Bootstrapping with 80 % of data 5 times
#> Completed
# retrieve hazard ratios
hazard_ratio(model)
#> Hazard ratio LCI UCI
#> 0.9582143 0.7107252 1.2918841Dose-response with 5 bootstrap samples and competing events in subgroups defined by sex
options <- SEQopts(km.curves = TRUE,
bootstrap = TRUE,
bootstrap.nboot = 5,
compevent = "LTFU",
# define the subgroup
subgroup = "sex")
data <- SEQdata.LTFU
model <- SEQuential(data, id.col = "ID",
time.col = "time",
eligible.col = "eligible",
treatment.col = "tx_init",
outcome.col = "outcome",
time_varying.cols = c("N", "L", "P"),
fixed.cols = "sex",
method = "dose-response",
options = options)
#> Non-required columns provided, pruning for efficiency
#> Pruned
#> Expanding Data...
#> Expansion Successful
#> Moving forward with dose-response analysis
#> Bootstrapping with 80 % of data 5 times
#> dose-response model created successfully
#> Creating Survival Curves for sex_0
#> Creating Survival Curves for sex_1
#> Completed
km_curve(model, plot.type = "risk")
#> $sex_0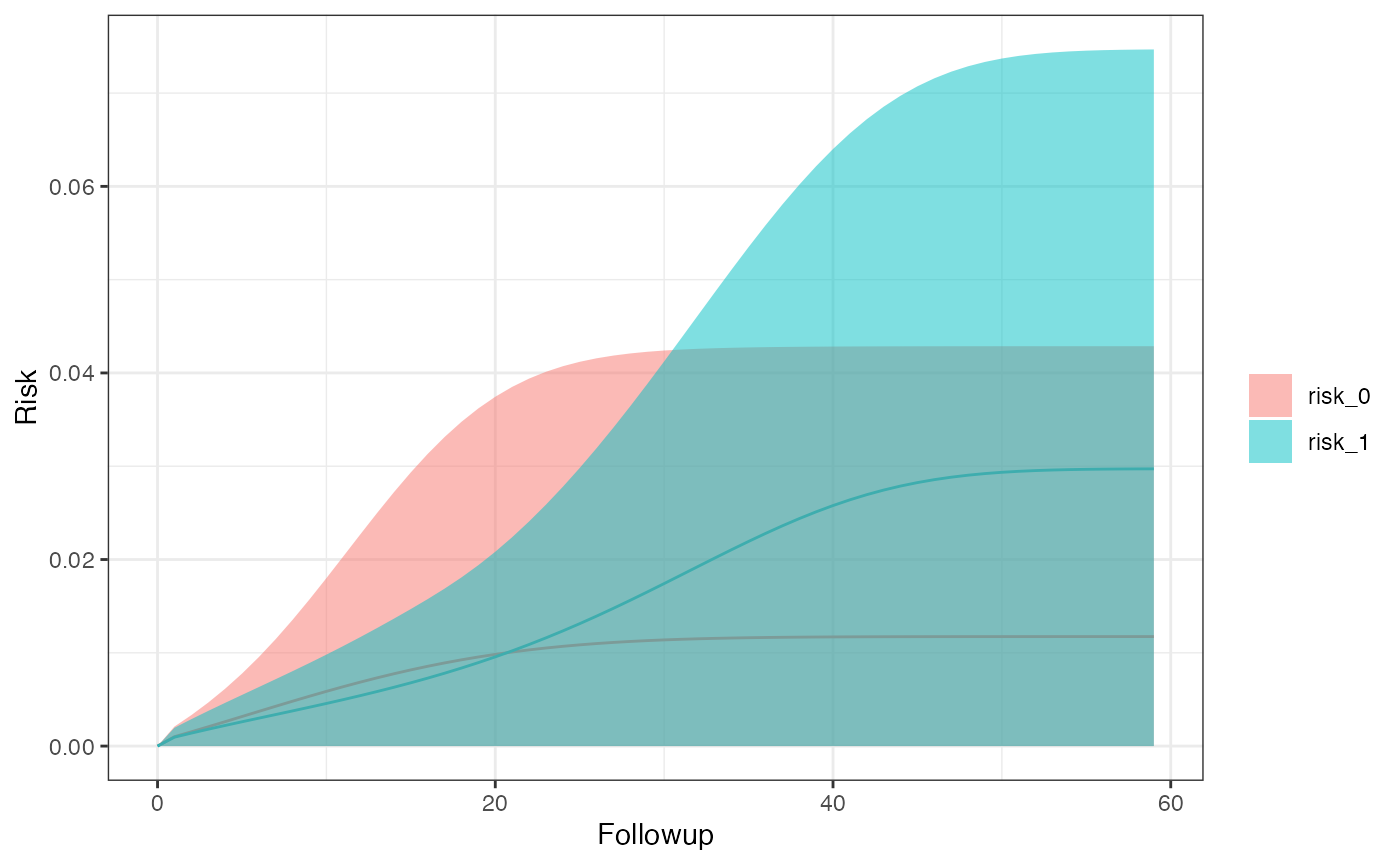
#>
#> $sex_1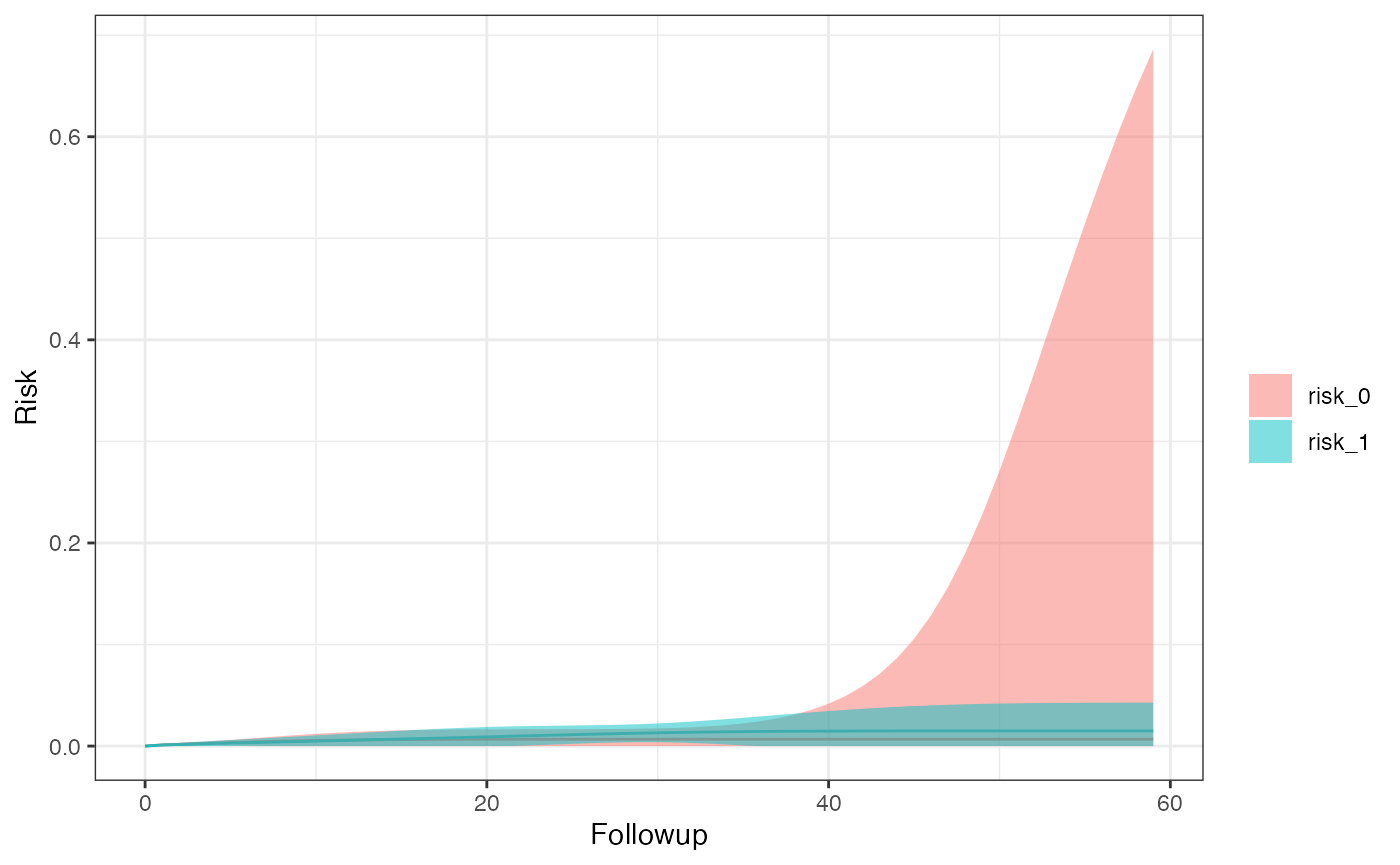
risk_data(model)
#> $sex_0
#> Method A Risk 95% LCI 95% UCI SE
#> <char> <char> <num> <num> <num> <num>
#> 1: dose-response 0 0.011257533 0 0.04037236 0.01485477
#> 2: dose-response 1 0.004588216 0 0.04979271 0.02306394
#>
#> $sex_1
#> Method A Risk 95% LCI 95% UCI SE
#> <char> <char> <num> <num> <num> <num>
#> 1: dose-response 0 0.006598380 0 0.47302574 0.23797751
#> 2: dose-response 1 0.004921131 0 0.03904126 0.01740855
risk_comparison(model)
#> $sex_0
#> A_x A_y Risk Ratio RR 95% LCI RR 95% UCI Risk Differerence
#> <fctr> <fctr> <num> <num> <num> <num>
#> 1: inc_0 inc_1 0.4075685 8.170464e-06 20330.8 -0.006669317
#> 2: inc_1 inc_0 2.4535753 4.918645e-05 122392.1 0.006669317
#> RD 95% LCI RD 95% UCI
#> <num> <num>
#> 1: -0.05854881 0.04521017
#> 2: -0.04521017 0.05854881
#>
#> $sex_1
#> A_x A_y Risk Ratio RR 95% LCI RR 95% UCI Risk Differerence RD 95% LCI
#> <fctr> <fctr> <num> <num> <num> <num> <num>
#> 1: inc_0 inc_1 0.745809 0.02567347 21.66559 -0.001677249 -0.4377930
#> 2: inc_1 inc_0 1.340826 0.04615613 38.95071 0.001677249 -0.4344385
#> RD 95% UCI
#> <num>
#> 1: 0.4344385
#> 2: 0.4377930